Understanding where species occur is a major challenge in biogeography, and of applied interest for forecasting species responses to climate change. How to best make these species distribution models has been a central discussion in recent decades. The idea underlying many models is Hutchinson’s n-dimensional hypervolume concept, but many models now in use do not now actually use this concept to make their predictions. So should hypervolumes be used for species distribution modeling applications?
A few years ago, we proposed a method to estimate n-dimensional hypervolumes based on kernel density estimation (Blonder et al. 2014). While the primary applications were in measuring niche geometry and functional diversity, we also showed an example of how the approach could be used to carry out species distribution modelling. The essence of the idea was to determine whether test points (e.g. different geographic locations) have climate axis values that correspond to regions within the kernel density estimate.
Recently this approach was critically discussed in the literature (Qiao et al. 2017). They cautioned against using hypervolumes based on kernel density estimates for niche modelling applications, especially in cases where fundamental niches were of interest. The central argument was that the approach had high error rates due to the ‘padding’ inherent to kernel density estimation – falsely predict species to be present when they were absent, and falsely predicting species to be absent when they were preset. They also suggested that simpler models (e.g. hyperboxes) were more appropriate for fundamental niches than the complex shapes that can arise from kernel density estimation.
The journal Global Ecology and Biogeography invited us to respond to these criticisms, and I am pleased that our formal response is now available online (Blonder et al. 2017 – PDF available here), jus now in the midst of a busy field season.
Writing this piece was a collegial and productive back-and-forth with the authors of the original criticism, and the process certainly helped clarify concepts for people on each side of the debate. Qiao et al. should have their own response to our response online shortly, but for now, I want to summarize our own arguments. have published their own response to our letter, which is well worth reading.
Briefly, we argue that the statistical performance of the kernel density estimation method is actually high, and comparable to many other leading methods. It is only when the assumptions of kernel density estimation (independent and identically distributed samples, i.e. no sampling bias) are not met that lower performance occurs. We also suggest that niche modelling methods that capture more complexity of the observed data are often useful. Indeed, many other niche modelling methods actually have reasonable interpretations as hypervolumes, as you can see below.
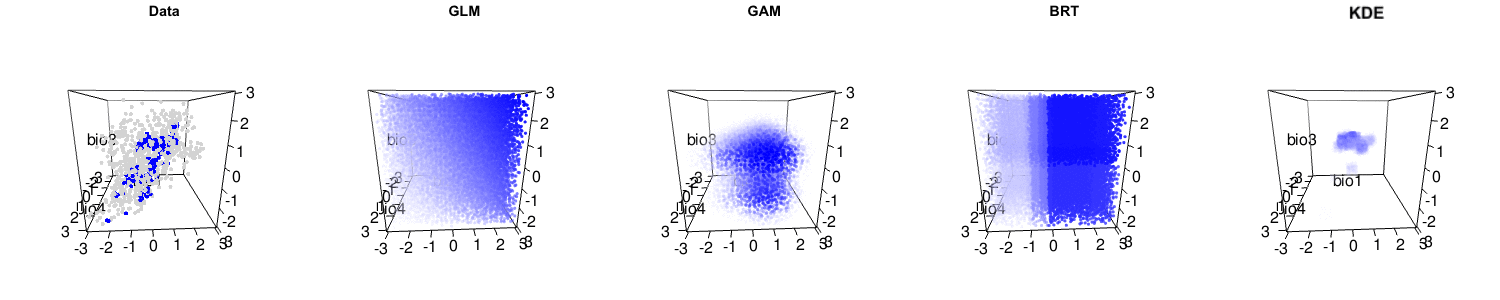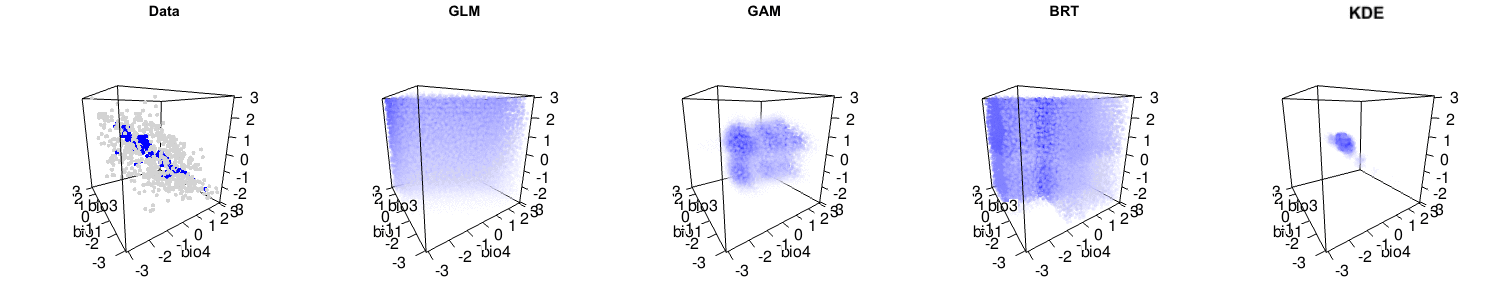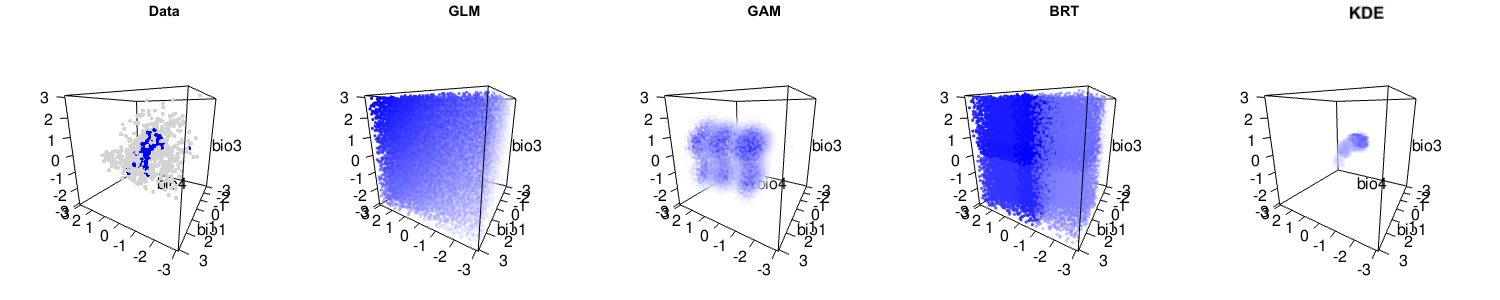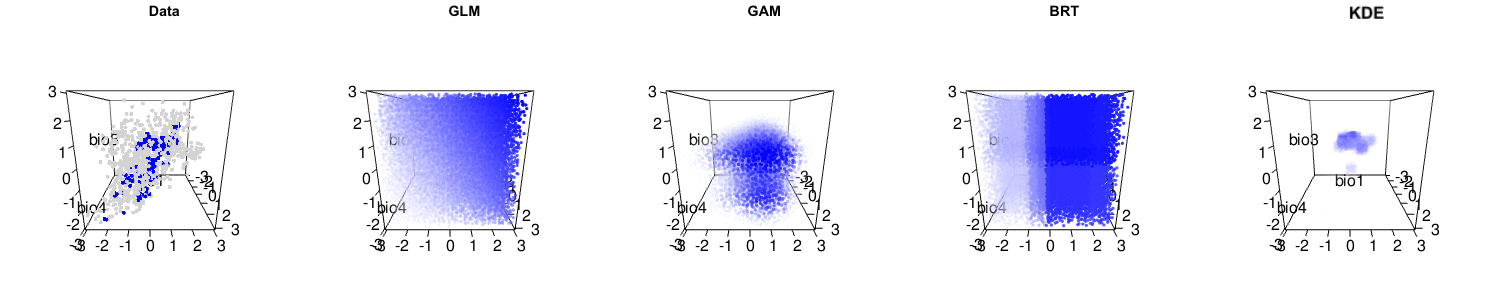
The debate is certainly far from over, and the central question of how to best model species’ distributions remains open. But please have a careful read of Qiao et al.‘s criticism and our response to make up your own mind about this important issue!


